Application of T2WI radiomics to predict 1p/19q status in low-grade gliomas
-
摘要:
目的 基于磁共振T2WI构建影像组学模型,预测低级别胶质瘤1p/19q缺失状态的价值。 方法 回顾性分析本院经病理证实的154例低级别胶质瘤患者(1p/19q共缺失100例,1p/19q非共缺失54例),按照分层抽样7∶3分成训练集和验证集。使用3D-Slicer软件对肿瘤区域进行手动分割,用pyradiomics进行特征提取。临床资料的分析采用t检验/χ2检验;影像组学特征采用方差法和10折交叉验证的LASSO算法进行筛选,最后建立支持向量机、高斯朴素贝叶斯、K-近邻、逻辑回归模型,采用ROC曲线的曲线下面积值和sklearn分类报告中的参考指标(准确度、敏感度、特异性、F1分数)进行效能评价。 结果 4种模型中,支持向量机的曲线下面积值最高,训练集和验证集分别为0.95、0.91;参考指标中表现最佳为K-近邻,其准确度、敏感度、特异性及F1分数分别为0.87、0.97、0.70、0.91;其次为支持向量机,各项指标与模型平均值相当。 结论 基于T2WI影像组学模型可以有效地预测低级别胶质瘤1p/19q的缺失状态。 Abstract:Objective To predict 1p/19q deletion status in low-grade gliomas by construction of the radiomics model based on magnetic resonance T2WI. Methods A total of 154 patients with low- grade gliomas confirmed by pathology in our hospital (including 100 cases of 1p/19q deletion and 54 cases of non-deletion of 1p/19q) were retrospectively analyzed and divided into training set and validation set according to stratified sampling at 7∶3. The tumor region was segmented manually by 3D-Slicer software and the features were extracted by pyradiomics. The clinical data were analyzed by t-test and χ2 test, and the radiomics features were screened by variance method and 10-fold cross- validation LASSO algorithm. Finally, the models of support vector machine, gaussian naïve bayes、K-nearest neighbor and logistic regreesion were established, and the efficacy was evaluated by the area under the ROC curve and the reference indexes (accuracy, sensitivity, specificity, F1 score) in sklearn classification report. Results Among the four models, the AUC value of support vector machine was the highest, the training set and verification set were 0.95 and 0.91 respectively, the best reference index was K-nearest neighbor, its accuracy, sensitivity, specificity and F1 score were 0.87, 0.97, 0.70, 0.91 respectively, followed by support vector machine, each index was equal to the average value of the model. Conclusion Based on the T2WI radiomics model, the state of 1p/19q deletion in low-grade gliomas can be effectively predicted. -
Key words:
- glioma /
- magnetic resonance imaging /
- radiomics /
- 1p/19q co-deletion
-
表 1 患者临床信息及染色体1p/19q共缺失状态
Table 1. Clinical information and chromosome 1p/19 co-deletion status [n(%)]
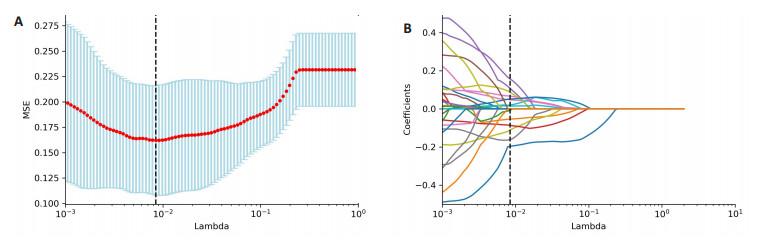
指标 1p/19q共缺失(n=100) 1p/19q非共缺失(n=54) 合计(n=154) 年龄(岁, Mean±SD) 43.28±13.03 39.85±14.39 42.08±13.58 性别 男 49(49.00) 32(59.26) 81(52.59) 女 51(51.00) 22(40.74) 73(47.41) WHO分级 2级 65(65.00) 37(68.52) 102(66.23) 3级 35(35.00) 17(31.48) 52(33.77) 病理学分类* 星形细胞瘤 - 54(100.00) 54(35.06) 少突胶质细胞瘤 100(100.00) - 100(64.94) *根据2021年WHO中枢神经系统肿瘤分类标准. 表 2 LASSO筛选的影像组学特征
Table 2. Radiomics features screened by LASSO algorithm
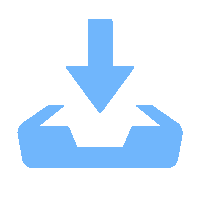
序号 特征名称 系数 feature29 original_firstorder_Skewness -0.1926 feature35 original_glcm_ClusterShade -0.0539 feature50 original_glcm_JointEnergy -0.0138 feature59 original_gldm_DependenceNonUniformityNormalized 0.1181 feature65 original_gldm_LargeDependenceHighGrayLevelEmphasis 0.0469 feature92 original_glszm_LargeAreaHighGrayLevelEmphasis -0.1671 feature100 original_glszm_ZoneEntropy -0.1189 feature146 wavelet-LLH_glcm_MaximumProbability 0.2001 feature185 wavelet-LLH_glszm_LargeAreaHighGrayLevelEmphasis 0.1774 feature215 wavelet-LHL_firstorder_Skewness 0.0645 feature246 wavelet-LHL_gldm_DependenceVariance 0.0729 feature392 wavelet-HLL_ firstorder_Kurtosis 0.0173 feature401 wavelet-HLL_firstorder_Skewness 0.0517 feature418 wavelet-HLL_glcm_Imc1 -0.0879 表 3 机器学习模型的效能评价指标
Table 3. Effectiveness evaluation index of machine learning models
模型 准确度 敏感度 特异性 F1分数 SVM 0.83 0.93 0.64 0.87 GNB 0.72 0.83 0.52 0.79 KNN 0.87 0.97 0.70 0.91 LR 0.83 0.90 0.70 0.87 平均* 0.81 0.91 0.64 0.86 *4种机器学习模型单一指标的均值. -
[1] Louis DN, Perry A, Wesseling P, et al. The 2021 WHO classification of tumors of the central nervous system: a summary[J]. Neuro Oncol, 2021, 23(8): 1231-51. doi: 10.1093/neuonc/noab106 [2] Anshit G. Erratum. The T2-FLAIR-mismatch sign as an imaging biomarker for IDH and 1p/19q status in diffuse low-grade gliomas: a systematic review with a Bayesian approach to evaluation of diagnostic test performance[J]. Neurosurg Focus, 2020, 48(5): E10. doi: 10.3171/2020.3.FOCUS19660a [3] Yogananda CGB, Shah BR, Yu FF, et al. A novel fully automated MRI-based deep-learning method for classification of 1p/19q co-deletion status in brain gliomas[J]. Neuro Oncol Adv, 2021, 2 (Supplement_4): iv42-8. [4] de Biase D, Acquaviva G, Visani M, et al. Next-generation sequencing panel for 1p/19q codeletion and IDH1-IDH2 mutational analysis uncovers mistaken overdiagnoses of 1p/19q codeletion by FISH[J]. J Mol Diagn, 2021, 23(9): 1185-94. doi: 10.1016/j.jmoldx.2021.06.004 [5] Riche M, Amelot A, Peyre M, et al. Complications after frame-based stereotactic brain biopsy: a systematic review[J]. Neurosurg Rev, 2021, 44(1): 301-7. doi: 10.1007/s10143-019-01234-w [6] Eckel-Passow JE, Lachance DH, Molinaro AM, et al. Glioma groups based on 1p/19q, IDH, and TERT promoter mutations in tumors[J]. N Engl J Med, 2015, 372(26): 2499-508. doi: 10.1056/NEJMoa1407279 [7] van Lent DI, van Baarsen KM, Snijders TJ, et al. Radiological differences between subtypes of WHO 2016 grade Ⅱ-Ⅲ gliomas: a systematic review and meta-analysis[J]. Neuro Oncol Adv, 2020, 2 (1): vdaa044. doi: 10.1093/noajnl/vdaa044 [8] 潘婷, 苏春秋, 张璇, 等. 常规MRI特征在预测弥漫性低级别胶质瘤1p/19q缺失状态的应用价值[J]. 临床放射学杂志, 2020, 39(6): 1189-94. https://www.cnki.com.cn/Article/CJFDTOTAL-LCFS202006036.htm [9] 樊建坤, 程勇, 王腾, 等. T2-FLAIR影像组学预测弥漫性较低级别胶质瘤1p/19q缺失状态的价值[J]. 中国医学影像学杂志, 2021, 29 (5): 425-9. doi: 10.3969/j.issn.1005-5185.2021.05.003 [10] Lasocki A, Gaillard F, Gorelik A, et al. MRI features can predict 1p/ 19q status in intracranial gliomas[J]. AJNR Am J Neuroradiol, 2018, 39(4): 687-92. doi: 10.3174/ajnr.A5572 [11] Fan ZW, Sun ZY, Fang SY, et al. Preoperative radiomics analysis of 1p/19q status in WHO grade Ⅱ gliomas[J]. Front Oncol, 2021, 11: 616740. doi: 10.3389/fonc.2021.616740 [12] Kong ZR, Jiang CD, Zhang YW, et al. Thin-slice magnetic resonance imaging-based radiomics signature predicts chromosomal 1p/19q Co-deletion status in grade Ⅱ and Ⅲ gliomas[J]. Front Neurol, 2020, 11: 551771. doi: 10.3389/fneur.2020.551771 [13] Kocak B, Durmaz ES, Ates E, et al. Radiogenomics of lower-grade gliomas: machine learning-based MRI texture analysis for predicting 1p/19q codeletion status[J]. Eur Radiol, 2020, 30(2): 877-86. doi: 10.1007/s00330-019-06492-2 [14] Sarvamangala DR, Kulkarni RV. Convolutional neural networks in medical image understanding: a survey[J]. Evol Intel, 2022, 15(1): 1-22. doi: 10.1007/s12065-020-00540-3 [15] Maximov Ⅱ, Vellmer S. Isotropically weighted intravoxel incoherent motion brain imaging at 7T[J]. Magn Reson Imaging, 2019, 57: 124-32. doi: 10.1016/j.mri.2018.11.007 [16] Nuessle NC, Behling F, Tabatabai G, et al. ADC-based stratification of molecular glioma subtypes using high b-value diffusion-weighted imaging[J]. J Clin Med, 2021, 10(16): 3451. doi: 10.3390/jcm10163451 -






 下载:
下载: