An aptamer-based analytical method for rapid detection of lung cancer tumor markers NSE antigen
-
摘要:
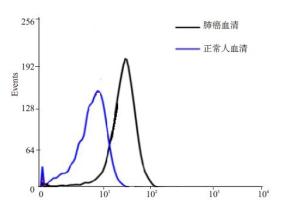
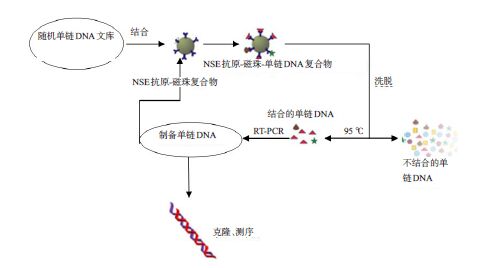
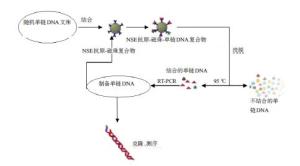
目的提出一种基于适配子快速检测肺癌肿瘤标志物的方法。 方法利用指数级富集配体的系统进化技术筛选其高特异性适配子,通过流式细胞术检测亲和力,最终选出一条高亲和力、强特异性的适配子。枸橼酸钠还原法制备纳米金颗粒。建立基于免标记纳米金和适配子的分析方法,并对其方法进行考察。 结果通过 6 轮筛选得到三条 NSE 的高特异性、强亲和力的适配子,其中,a 适配子特异性最高。同时利用筛选得到的肺癌肿瘤标志物 a 适配子和纳米金颗粒建立一种快速检测 NSE 抗原的方法。 结论该方法具有较好的准确度,为肺癌的早期检测提供了一种新的手段。 -
关键词:
- 神经元特异性烯醇化酶抗原 /
- 适配子 /
- 纳米金 /
- 肺癌 /
- 肿瘤标志物
Abstract:ObjectiveTo explore a new aptamer-based rapid detection method for lung cancer tumor markers. MethodsHighly specific adaptive were screened by using systematic evolution of ligands (SELEX) and affinity were detected by flow cytometry. So as to choose a high affinity and high specificity of aptamer. Gold nanoparticles were prepared through sodium citrate reduction. The method based on label-free analysis of gold nanoparticles and aptamers were established, with methodological investigation. ResultsHigh specificity and strong affinity of 3 NSE were obtained through 6 rounds of screening, with highest specificity of a. A rapid method for the detection of NSE antigen were established through the selection of tumor markers of lung cancer and gold nanoparticles. ConclusionThe method has a good accuracy for the early detection of lung cancer provides a new means. -
Key words:
- NSE antigen /
- aptamers /
- nanometer gold /
- lung cancer /
- tumor markers
-
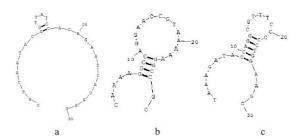
表 1 适配子测序序列
序列号 大小(bp) 序列 Kd ( mmol/L) a 30 CGGTAATACGGTTATCCACAGAATCAGGGG 21±8 b 29 CAAAAGGCCAGGAACCGTAAAAAGGCCGC 30±5 c 30 TAAAGATACCAGGCGTTTCCCCCTGGAAGC 33±4 -
[1] Cai D, Xu D, Zhang Q, et al. Classification of lung cancer using ensemble-based feature selection and machine learning methods[J]. Molecular biolystems, 2014, 464(7291): 1071-6. http://cn.bing.com/academic/profile?id=49fee408729c733e13f837e4fc1d38b7&encoded=0&v=paper_preview&mkt=zh-cn [2] 杜淑英. 血清5种肿瘤标志物联合检测在肺癌诊断中的价值[J]. 北方药学, 2014, 11(9): 85-6. http://www.cnki.com.cn/Article/CJFDTOTAL-BFYX201409073.htm [3] 时广利, 胡秀玲, 岳思东, 等. 血清肿瘤标志物在肺癌辅助诊断中的应用[J]. 中华肿瘤杂志, 2005, 27(5): 299-301. http://www.cnki.com.cn/Article/CJFDTOTAL-ZHZL200505014.htm [4] Mckeague M, Mcconnell EM, Cruztoledo TJ, et al. Analysis of in vitro aptamer selection parameters[J]. J Mole Evol, 2015,81(5):150-61. http://cn.bing.com/academic/profile?id=cde2fce0e2162906faf59b21657e4d47&encoded=0&v=paper_preview&mkt=zh-cn [5] Liu K, Lin B, Lan X. Aptamers: a promising tool for cancer imaging, diagnosis, and therapy[. J]. J Cell Biochem, 2013, 114(2): 250-5. doi: 10.1002/jcb.v114.2 [6] Darmostuk M, Rimpelová S, Gbelcova H, et al. Current approaches in SELEX: An update to aptamer selection technology.[J]. Biotechnol Adv, 2015, 33(6): 1141-61. doi: 10.1016/j.biotechadv.2015.02.008 [7] Chen X, Zhang J, Nilsen HM. Fe-g-C3N4-Catalyzed oxidation of benzene to phenol using hydrogen peroxide and visible light[J]. J Am Chem Soc, 2009, 131(33): 12018-24. http://cn.bing.com/academic/profile?id=f291bdceaf085dc88a6cdece58616676&encoded=0&v=paper_preview&mkt=zh-cn [8] 陈伶利, 李杰, 张晓青, 等. 以大肠埃希菌tolC蛋白为靶标的适配体的筛选及结构分析[J]. 北京大学学报: 医学版, 2014, 46(5): 698-702. http://www.cnki.com.cn/Article/CJFDTOTAL-BYDB201405006.htm [9] Fletcher J, Phillips W, Milligan S, et al. Toward specific detection of dengue virus serotypes using a novel modular biosensor[J]. Biosens Bioelectron, 2010, 26(4): 1696-700. doi: 10.1016/j.bios.2010.07.046 [10] Biroccio A, Hamm J, Incitti I, et al. Selection of RNA aptamers that are specific and high-affinity ligands of the hepatitis C virus RNA-dependent RNA polymerase[J]. J Virol, 2002, 76(8): 3688-96. doi: 10.1128/JVI.76.8.3688-3696.2002 [11] Zhou J, Shu Y, Guo P, et al. Dual functional RNA nanoparticles-gp120 aptamer for cell-type specific delivery and HIV-1 inhibition[J]. Mrthods, 2011, 54(2): 284-94. [12] Wheeler A, Trifonova R, Vrbanac V, et al. Inhibition of HIV transmission in human cervicovaginal explants and humanized mice using CD4 aptamer-siRNA chimeras[J]. J Clin Invest, 2011,121(6): 2401-12. doi: 10.1172/JCI45876 [13] Tuerk C, Gold L. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase[J]. Science, 1990, 249(4968): 505-10. doi: 10.1126/science.2200121 [14] Fang XH, Sen A, Vicens M, et al. Synthetic DNA aptamers to detect protein molecular variants in a high-throughput fluorescence quenching assay[J]. Chembiochem, 2003, 4(9): 829-34. doi: 10.1002/cbic.v4:9 [15] Jeon H, Kayhan B, Ben YT, et al. A DNA aptamer prevents influenza infection by blocking the receptor binding region of the viral hemagglutinin[J]. J Biol Chem, 2004, 279(46): 48410-9. doi: 10.1074/jbc.M409059200 [16] Misono S, Kumar K. Selection of RNA aptamers against human influenza virus hemagglutinin using surface plasmon resonance[J]. Anal Biochem, 2005, 342(2): 312-7. doi: 10.1016/j.ab.2005.04.013 [17] Gopinath C, Misono S, Kawasaki K, et al. An RNA aptamer that distinguishes between closely related human influenza viruses and inhibits haemagglutinin-mediated membrane fusion[J]. J Gen Virol,2006, 87(Pt 3): 479-87. http://cn.bing.com/academic/profile?id=72070702d4e4646526b119406a48a96c&encoded=0&v=paper_preview&mkt=zh-cn -





 下载:
下载: