Multi gene promotion in hepatocellular carcinoma
-
摘要:
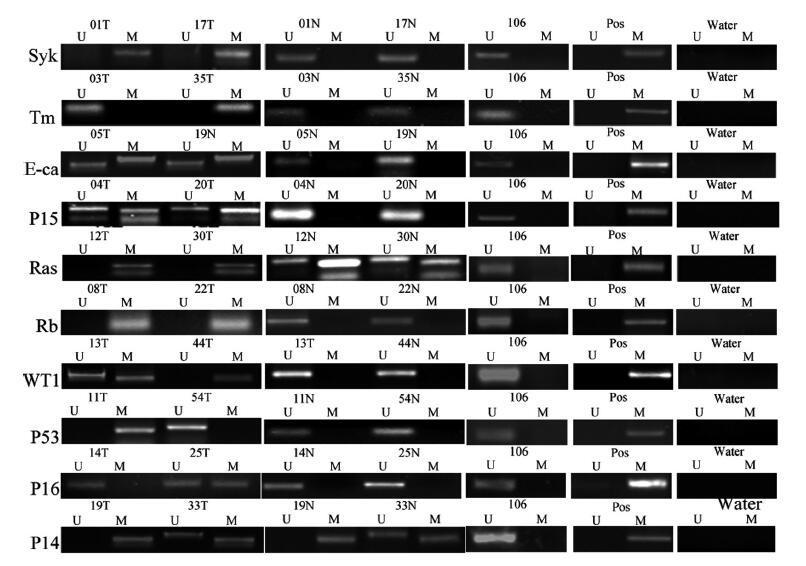
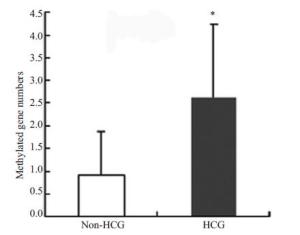
目的 通过研究肝癌中多基因启动子区的甲基化,说明抑癌基因甲基化在肝癌发生的普遍性。 方法分别抽提冻存的肝癌、癌旁组织和正常肝组织的DNA,利用亚硫酸氢钠具有使未甲基化的胞嘧啶转为尿嘧啶而不改变甲基化胞嘧啶的特性分别设计甲基化引物和非甲基化引物,然后分别进行PCR 反应。我们分别检测了转移抑制基因p15, SYK, TIMP-3, E-cadherin,RASSAF1 和肿瘤相关基因p53, RB1, WT1, p14, p16启动子区在60个肝癌、癌旁和6个正常肝组织的甲基化情况。 结果60个肝癌标本中10个基因的甲基化情况各不相同:从p53的8%~90%的RASSAF1;肝癌组织比癌旁组织的甲基化平均水平高,二者在统计学上有显著差异(P<0.05)。 结论肝癌中有多个基因存在甲基化,且肝癌组织甲基化水平较癌旁组织高。 Abstract:Objective To explain the aberrant methylation of tumor suppressor genes is common hepatocellular carcinoma. MethodsExtracting cryopreservation of hepatocellular carcinoma, paracancerous tissues and normal liver tissues of DNA,using sodium bisulfite has the characteristics of unmethylated cytosine to uracil without changing the methylated cytosine methylation primers were designed and the non methylated primers, and then respectively by PCR reaction. We examined the metastasis suppressor gene p15, SYK, TIMP-3, E-cadherin, RASSAF1 and tumor related genes p53, RB1, WT1, P14, p16 promoter methylation status of the promoter region in 60 liver cancer, para cancer and 6 normal liver tissues. ResultsThefrequency of promoter methylation of ten genes in 60 HCC varied from 8% in P53 to 90% in p14. The average methylated-gene numbers were significantly different between HCC and paired non-tumor tissues (P < 0.05). Conclusion There are multiple gene methylation in HCC exist, and the level of methylation of liver cancer tissue than in noncancerous tissue high. -
Key words:
- hepatocellular carcinoma /
- gene /
- methylation
-
表 1 基因甲基化和未甲基化引物
基因 引物序列(5'-3') 产物(bp) 参考文献 U F* ATTTTGTGGGTTTTGTTTGGTG 140 SYK R ACTTCCTTAACACACCCAAAC M F CGATTTCGCGGGTTTCGTTC 243 R AAAACGAACGCAACGCGAAAC U F TGTGATGTGTTTGTATTTTGTGGTT 154 P15 R CCATACAATAACCAAACAACCAA M F GCGTTCGTATTTTGCGGTT 148 R CGTACAATAACCGAACGACCGA U F TGGTTGTAGTTATGTATTTATTTTTAGTGGTGTT 120 E-cadherin R ACACCAAATACAATCAAATCAAACCAAA M F TGTAGTTACGTATTTATTTTTAGTGGCGTC 112 R CGAATACGATCGAATCGAACCG U F TTTTGTTTTGTTATTTTTTGTTTTTGGTTTT 122 TIMP-3 R CCCCCAAAAACCCCACCTCA [5] M F CGTTTCGTTATTTTTTGTTTTCGGTTTC 116 R CCGAAAACCCCGCCTCG U F TTTGGTTGGAGTGTGTTAATGTG 105 RASSF1A R CAAACCCCACAAACTAAAAACAA [5] M F GTGTTAACGCGTTGCGTATC 93 R AACCCCGCGAACTAAAAACGA U F TTATTAGAGGGTGGGGTGGATTGT 151 P16 R CAACCCCAAACCACAACCATAA [5] M F TTATTAGAGGGTGGGGCGGATCGC 150 R GACCCCGAACCGCGACCGTAA U F TTGGTAGGTGGATTATTTGTTT 247 P53 R CCAATCCAAAAAAACATATCAC [5] M F TTCGGTAGGCGGATTATTTG 193 R AAATATCCCCGAAACCCAAC U F TGGGATTTGGGTGGTATTTG 124 WT1 R CACCAACACCCACTACACCA [5] M F GTTAGGCGTCGTCGAGGTTA 110 R AAAACGCAAAATCCAACACC U F TTTTTGGTGTTAAAGGGTGGTGTAGT 132 P14 R CACAAAAACCCTCACTCACAACAA [5] M F GTGTTAAAGGGCGGCGTAGC 122 R AAAACCCTCACTCGCGACGA U F GGGAGTTTTGTGGATGTGAT 172 RB1 R ACATCAAAACACACCCCA [5] M F GGGAGTTTCGCGGACGTGAC 172 R ACGTCGAAACACGCCCCG M,甲基化;U,未甲基化. -
[1] El-Serag HB. Hepatocellular carcinoma -An epidemiologic view [J]. J Clin Gastroenterol, 2002, 35(5, 2): S72-8. [2] 熊兰, 曾斌. DNA甲基化与肝癌关系研究进展[J]. 社区医学杂志,2013, 11(19): 24-6. [3] 胡静娴, 谢渭芬. DNA甲基化与肝癌的研究现状[J]. 国际消化病杂志, 2013, 33(1): 25-8. [4] 孙其恺, 王伟, 荚卫东. 表观遗传修饰与肝癌的关系[J]. 中华消化外科杂志, 2013, 12(9): 717-20. [5] Jones PA, Baylin SB. The fundamental role of epigenetic events in cancer[J]. Nat Rev Genet, 2002, 3(6): 415-28. [6] Davis CD, Uthus EO. DNA methylation,Cancer susceptibility,and nutrient interactions[J]. Exp Biol Med, 2004, 229(2): 988-95. [7] Caiafa P, Zampieri M. DNA methylation and chromatin structure: The puzzling CpG islands[J]. J Cell Biochem, 2005, 94(2): 257-65. [8] Sidransky D. Emerging molecular markers of cancer[J]. Nat Rev Cancer, 2002, 2(3): 210-9. -






 下载:
下载: